There are 20,000 “genes” but more than 100,000 different proteins.
There are 20,000 “genes” but more than 100,000 different proteins.
Alternative RNA editing is necessary to form the many different patterns.
In the fly, one gene has 38,000 alternative patterns. (picture below)
In humans, one gene can have 500 alternative patterns. (picture below)
Alternative RNA splicing in evolution is one of the critical factors in the development of the complexity of human beings
 Previous posts have shown how the shapes of complex interlocking proteins allow the neuron machinery to process mental events. The shapes of the proteins are determined by the DNA code, which determines the amino acids of the protein, the folding and eventual protein shape. But, the determination of the final code that is used is not so simple. As important as the DNA code is, the alternative RNA splicing is equally, or more, important.
Previous posts have shown how the shapes of complex interlocking proteins allow the neuron machinery to process mental events. The shapes of the proteins are determined by the DNA code, which determines the amino acids of the protein, the folding and eventual protein shape. But, the determination of the final code that is used is not so simple. As important as the DNA code is, the alternative RNA splicing is equally, or more, important.
In fact, recent research shows that alternative splicing may be the critical source of evolutionary changes differentiating primates and humans from other creatures such as worms and flies with a similar number of genes.
Dogma, Old and New
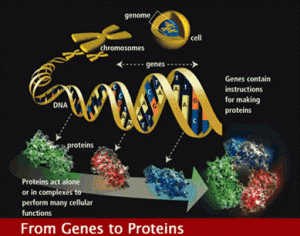 The old genome dogma was simple, DNA copies onto messenger RNA, which then travelled to the ribosome where proteins were made. But, the cell didn’t oblige with this dogma. It is not that simple.
The old genome dogma was simple, DNA copies onto messenger RNA, which then travelled to the ribosome where proteins were made. But, the cell didn’t oblige with this dogma. It is not that simple.
Years ago it became clear that when DNA was copied into RNA, many sections that were not translated into the protein, called introns, had to be edited out. Later, the genome project showed that there are only 20,000 genes identified, representing a tiny 1.5% percentage of the DNA in the chromosome. The rest was considered “junk”, unused, useless DNA.
This was very disappointing because it was known that there are perhaps at least 100,000 different types of proteins in the human body, and probably many more. Also many of the proteins in humans have subtle differences from other creatures. How could so few genes make so many different proteins?
The answer to this problem was discovered with alternative splicing of the RNA messenger transcript before it goes to the ribosome to make proteins.
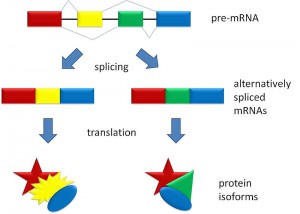 In eukaryotes, all animals and plants, the code of the DNA strand is copied into a RNA strand, that is a draft of what will later be used to build the protein. The messenger RNA will be transported to the ribosome where each specific amino acid is brought to the growing protein strand.
In eukaryotes, all animals and plants, the code of the DNA strand is copied into a RNA strand, that is a draft of what will later be used to build the protein. The messenger RNA will be transported to the ribosome where each specific amino acid is brought to the growing protein strand.
But, before the messenger RNA is finalized a “pre-messenger RNA” (pre-mRNA) is formed which includes sections of DNA that will be used for the protein code, called exons, and usually longer sections that will be edited out, called introns.
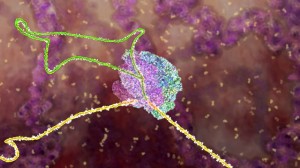
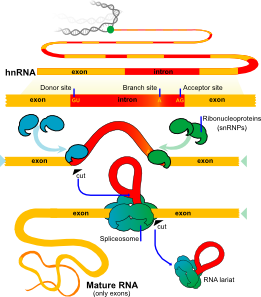 In many cases, an elaborate molecular machine, called the spiceosome, accomplishes this critical editing. The spliceosome is composed of multiple pieces of small nuclear RNA (snRNA) named U1, U2, U4, U5, and U6, with accompanying protein, (small nuclear RNA protein = snRNP). It is this very large complex machinery made from multiple RNA pieces and proteins that does the actual splicing. How this is directed is not so clear.
In many cases, an elaborate molecular machine, called the spiceosome, accomplishes this critical editing. The spliceosome is composed of multiple pieces of small nuclear RNA (snRNA) named U1, U2, U4, U5, and U6, with accompanying protein, (small nuclear RNA protein = snRNP). It is this very large complex machinery made from multiple RNA pieces and proteins that does the actual splicing. How this is directed is not so clear.
Nothing is simple. Splicing can also occur by other methods, such as the RNA strand forming a ribozyme, that is an RNA that acts as an enzyme to cut the strand. This is called self-splicing. Another technique involves the transfer RNA.
The So Called Gene
A previous post noted that after ENCODE (Encyclopedia of DNA Elements – see post 1, post 2, and post 3) some experts referred to a “so called gene.” This is because the part of the DNA that was once thought to be the code that makes protein is not really the final determinant of the code. In fact it was shown that the messenger RNA could pull DNA from multiple supposed genes, greatly muddying the concept of “gene”.
Although alternative splicing was known for many years, it was considered that exons and introns were fixed, and the splicing processes merely identified which DNA are exons and which were introns, and the introns were edited out of the transcript. This process was thought to be based upon specific code factors with some signaling where the cuts will appear. Also, regulatory particles, both proteins and microRNA were involved.
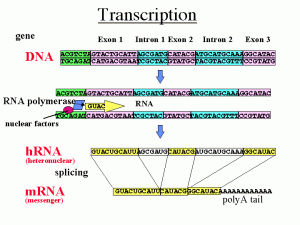 What has been recently discovered is that in different types of cells, the very same sequence of DNA can be considered an exon or an intron, creating a tremendous variation in the construction of proteins between different cells. Also, the points at which the cut is made is not as fixed as it seemed at first. Therefore, it has been found that specific types of cells, such as those from organs like the brain, the kidney, etc., have multiple alternative splicing patterns.
What has been recently discovered is that in different types of cells, the very same sequence of DNA can be considered an exon or an intron, creating a tremendous variation in the construction of proteins between different cells. Also, the points at which the cut is made is not as fixed as it seemed at first. Therefore, it has been found that specific types of cells, such as those from organs like the brain, the kidney, etc., have multiple alternative splicing patterns.
The number of different protein patterns can be 2 up to 38,000 in the fly (which is more than their 14,000 genes). In human beings there are examples of 500 different patterns for one gene. (see pictures)
Specific cells have self-editing mechanisms that can determine their own specific protein shapes, and machinery.
Types of Splicing
There are many different types of messenger RNA splicing, some types have been clearly defined, but continued research seems to come up with more and more different kinds.
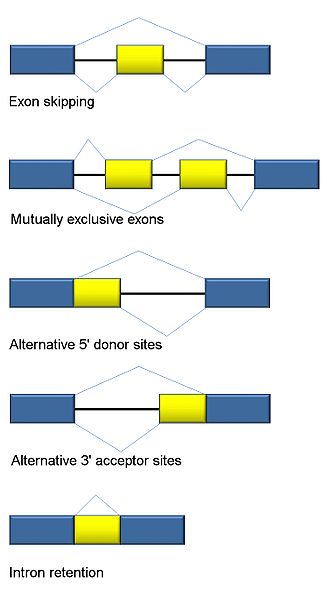 The first type involves introns being cut out.
The first type involves introns being cut out.
A second involves an exon (usually considered a piece that will stay) being spliced out.
Another involves two exons in a series, with only one of the two being kept in an alternating pattern.
There are a number of types of alternative splicing patterns where different parts of the intron are cut. Sometimes, mutations near the edge of the intron can affect this type of process.
Another type is where the intron is retained.
But, in fact, there are a large number of different types and the number is growing. This is also complicated by the fact that there can be multiple different steps in the process.
Splicing Regulation
There can be as many as 100 different introns and exons in a pre messenger RNA strand.
As mentioned above, some of the regulatory particles are proteins that come either from a nearby strand such as the DNA or RNA being utilized for the protein, or one that is produced at a very distant location, perhaps another RNA or a totally different piece of DNA. At first it was believed that there had to be a limited standard number of proteins, or the regulation would be have to be extremely vast, much more than many genomes could provide.
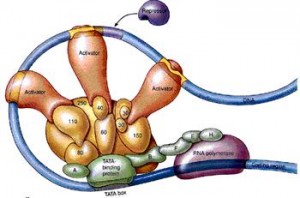 The proteins from distant locations are called “trans” regulatory particles, which are termed repressors or activators. Those proteins that are produced nearby, possibly on the same RNA strand, are called “cis” regulatory particles and termed silencers and enhancers. A silencer will stop the cut nearby, and the enhancer will bring about a cut nearby. But, in fact these distinctions of repressors and activators has broken down since there are now examples where all activators have functioned as repressors and all repressors as activators.
The proteins from distant locations are called “trans” regulatory particles, which are termed repressors or activators. Those proteins that are produced nearby, possibly on the same RNA strand, are called “cis” regulatory particles and termed silencers and enhancers. A silencer will stop the cut nearby, and the enhancer will bring about a cut nearby. But, in fact these distinctions of repressors and activators has broken down since there are now examples where all activators have functioned as repressors and all repressors as activators.
As with the genetic regulation that we observed in ENCODE there are a very large number of possible mechanisms. These different factors appear to work together to bring about a complicated outcome.
The Splicing Code
The ways that RNA splices occur has mushroomed in complexity, like other regulatory processes. At first the simple modes above were considered common. Now, research shows that there are hundreds of characteristics of RNA structures, and regulatory particles, which can be protein or microRNA, which determine how these alternative splicings occur.
For example, it used to be thought that the codes at the edges of the intron are most relevant. Now it appears that structures deep in the intron away from the cut may be quite relevant. There are dramatic differences in splicing triggered during the different phases of embryonic development. Many differences occur among different types of tissue cells. Other factors that affect splicing patterns include transcription rates, core-splicing-machinery levels, competition between splice sites, chromatin structure affecting the rate of transcription, histone modifications, and local chromatin modifications.
Recent articles attempting to define a splicing code have found that the regulation of this many different variables could take DNA directions the size of many additional genomes. It has been found in many cases of splicing there are multiple steps, which makes matters much more complex.
The number of variables is now so great that it has become another vast computation problem similar to the folding problem of the protein, and genome regulation in ENCODE. Please see the post on protein folding where it was noted that at a billion folds per second it would take ten billion years to try all possible folds of an average sized protein.
Layers upon Layers of Regulation
There are now at least three overlapping problems whose calculation is much greater than all of the fastest and greatest supercomputers put together could solve. In fact they are far beyond current capacities.
Attempts are now being made to conceptualize how these problems might fit into a large network.
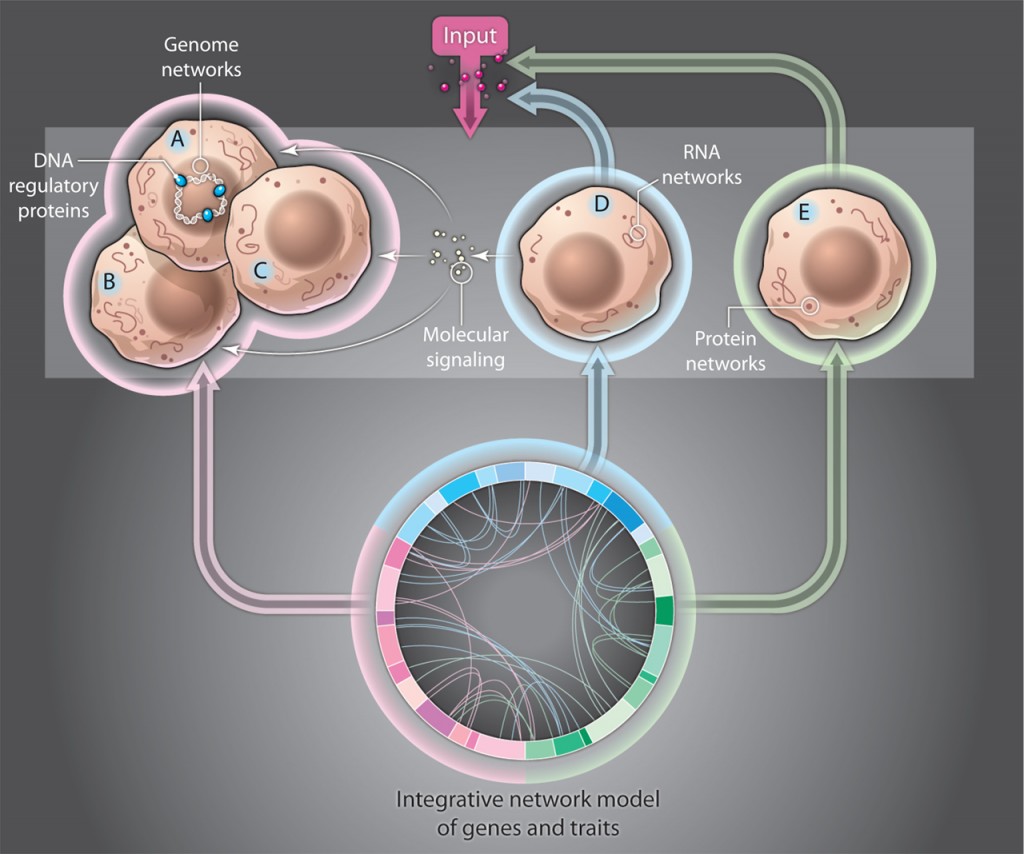
How does regulation occur with millions of regulatory particles and locations? This problem is addressed in the posts on ENCODE (post 1, post 2, post 3).
How does protein folding occur? See post on Folding and Protein shapes in the neuron.
How do the enormous amount of factors in the splicing code determine the new protein?
Alternative RNA Splicing in Evolution
Perhaps the most important recent finding is that alternative splicing is very different in different species. Specifically, the primates, and especially, the human being have by far the most complex alternative splicing. It now appears that alternative splicing is, perhaps, the most critical evolutionary factor determining the differences between human beings and other creatures.
A recent study showed that specific genes are activated in different types of cells, such as the different tissues, kidney, heart and brain. This type of gene activation seems to be similar in many different species (conserved is the evolutionary word.) These genes demonstrate very slow evolutionary changes.
But, with worms and flies having a similar number of genes as humans, there has to be more to explain the great differences. It now appears that the differences are in the alternative splicing. (Picture below is a human gene that has 500 alternative splicings, and a fly gene that has 38,000 alternative splicings, much more than the number of total genes in the fly)
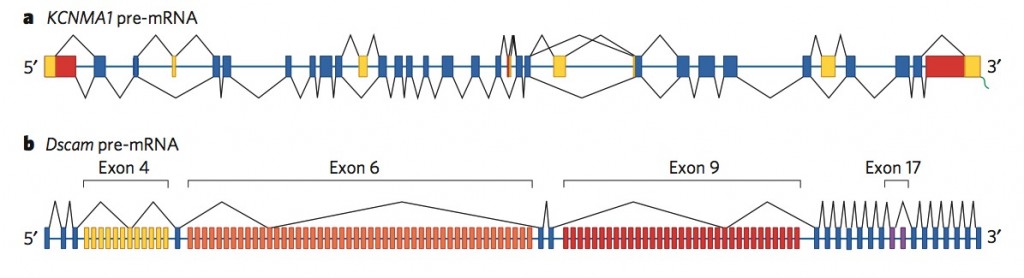
b. Fly pre mRNA up to 38,000 alternatives – blue regular – yellow, orange, red and purple alternatives
This new data shows that the uniqueness of the primates and humans is not in the specific genes triggered in the specific tissues, but a whole host of unique alternative splicing in all of the human cells, including unique types in each of the tissues. Also these alternative splicing changes occur rapidly in evolution.
Changing of genes is a very slow process that occurs over many millions of years. Changing of alternative splicing can happen in evolution much faster.
Different species have different alternative splicing and also have different diseases. There now appear to be the possibility that unique human diseases might be best understood in the unique alternative splicing of the human beings.
Evolution of the Human Being – Rewiring Human Signals
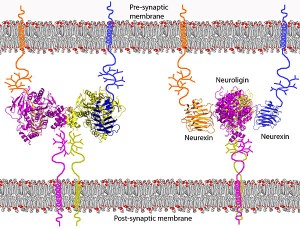 Human alternative splicing is being found to be critical throughout the body, but perhaps most important in the brain. In a previous post on proteins in the neuron, it was shown how critical the cell adhesion molecules are for synapses. Recent research has shown that alternative splicing modulates the interactions between the neuronal synaptic cell-adhesion molecules neuroligin 1 and neurexins, critical adhesion molecules holding the synapse in shape, connecting the pre and post synaptic neuron and the extracellular matrix.
Human alternative splicing is being found to be critical throughout the body, but perhaps most important in the brain. In a previous post on proteins in the neuron, it was shown how critical the cell adhesion molecules are for synapses. Recent research has shown that alternative splicing modulates the interactions between the neuronal synaptic cell-adhesion molecules neuroligin 1 and neurexins, critical adhesion molecules holding the synapse in shape, connecting the pre and post synaptic neuron and the extracellular matrix.
A very important example of alternative splicing involves the use of phosphorylation, that is, adding high-energy phosphorus onto a molecule. This process is critical to many important signaling channels in cells, and especially neurons. The proteins that perform phosphorylation are called kinases, and are the largest and most critical protein family for all types of cellular functions. They are the mechanism of a large number of the most important neurotransmitters and receptors in the brain.
 Phosphorylation triggers changes in a cascade of molecules called second messengers that take signals from receptors in the membranes all the way to the nucleus. Phosphorylation drives metabolism.
Phosphorylation triggers changes in a cascade of molecules called second messengers that take signals from receptors in the membranes all the way to the nucleus. Phosphorylation drives metabolism.
The many varied kinases, through advanced alternative splicing, determines that the human being has a much more complex system of signaling.
Alternative splicing affects the proteins of gene regulation, and all protein-protein interactions. Basically, alternative splicing affects all bodily functions in the human being.
Because alternative splicing is very significant for determining regulatory proteins, the regulation of alternative splicing piggy backs on the regulation of gene protein interactions, making it all even more complex.
The signaling changes from alternative splicing affect all human bodily functions. It is like rewiring the signaling outputs. In humans this rewiring is much more complex, perhaps the vast difference we see between humans and other animals.
Unique Human Protein Shapes
At first it appeared that RNA splicing involved editing out introns. Cuts appeared to be related to specific codes at the edges of the introns. Then it appeared that there were many different types of edits. Later, at times, introns and exons changed places. Cuts appeared in places where they shouldn’t. Splicing involved many different steps in a sequence. Histones, and chromatin patterns became involved.
The attempted splicing code became as complex as the protein folding code, that is, too complex to figure out.
 ENCODE showed that there are millions of regulatory particles, at least 18,000 small and large RNA particles. These newly discovered microRNA added to the enhancers and repressors of old.
ENCODE showed that there are millions of regulatory particles, at least 18,000 small and large RNA particles. These newly discovered microRNA added to the enhancers and repressors of old.
Perhaps, most shocking of all, ENCODE found that to make one protein RNA can take DNA code from multiple different areas that were once called multiple different genes. These different strands of DNA from different “genes” are edited together to form one new protein. At times, the old genes seem to expand and contract.
The interlocking systems of DNA regulation, RNA editing, and protein folding are what determine the interlocking shapes of the proteins in the neuron. These shapes form the machinery for mental events in the neuron.
 Alternative splicing is most advanced and rapidly developed in the evolution of the human being. In fact, alternative RNA splicing appears to be a strong determinant of the evolution of the human being.
Alternative splicing is most advanced and rapidly developed in the evolution of the human being. In fact, alternative RNA splicing appears to be a strong determinant of the evolution of the human being.
Where is the regulation for all of this? What determines what proteins are needed and what shapes they will be, the shapes needed for interlocking complex processes?