 How can microbes behave as if they have a brain? Microbes demonstrate complex decision-making and send specific chemical signals to colony members and other species, such as plants and animals. Their signals trigger group actions and behavioral changes including attacks. Slime mold microbe colonies show memory. Individual social amoebas join together and behave as a multicellular creature, then, divide again into individual cells. Individual microbes remember where they have travelled and can even plan for the future. In one very complex scenario, microbes start a multi step process to form a spore to avoid starvation, while continuing to make calculations about whether it is necessary. It is clear that these one-celled creatures can make decisions based upon many different sensory inputs. How can a cell behave as if it has a brain? How do microbes make decisions?
How can microbes behave as if they have a brain? Microbes demonstrate complex decision-making and send specific chemical signals to colony members and other species, such as plants and animals. Their signals trigger group actions and behavioral changes including attacks. Slime mold microbe colonies show memory. Individual social amoebas join together and behave as a multicellular creature, then, divide again into individual cells. Individual microbes remember where they have travelled and can even plan for the future. In one very complex scenario, microbes start a multi step process to form a spore to avoid starvation, while continuing to make calculations about whether it is necessary. It is clear that these one-celled creatures can make decisions based upon many different sensory inputs. How can a cell behave as if it has a brain? How do microbes make decisions?
Recent research has begun to shed some light on the very unusual abilities of these single cells. While study of metabolism is extremely complex, scientists are beginning to understand that microbe decision-making can be based on molecules in metabolic cycles that function as information signals in complex circuits of their “brain.”
Mind With No Brain
 In a previous post, Mind with No Brain, six features of microbes’ cognitive abilities were listed that are similar to brains. Microbes receive sensory information, integrate and analyze it and take appropriate actions. Microbes are able to do this without neurons. They solve problems such as finding food, evading predators, and initiate complex group activity.
In a previous post, Mind with No Brain, six features of microbes’ cognitive abilities were listed that are similar to brains. Microbes receive sensory information, integrate and analyze it and take appropriate actions. Microbes are able to do this without neurons. They solve problems such as finding food, evading predators, and initiate complex group activity.
- Microbes have specialized sensors over much of their membrane, which form a specialized honeycomb lattice with extreme chemical sensitivity. Thousands of protein receptors trigger responses. These receptors can sense the chemicals in the environment as small as 1 part in 1,000 and can trigger linking proteins and enzyme cascades for responses.
- Cilia can send electrical signals from one part of the microbe to another. If it hits a wall, the current travels to other cell parts and changes the direction and beat of the cilia. It is analogous to neuron communication.
- Microbes use proteins that contract and interact, similar to neurons during migration.
 Microbes secrete specific signals to communicate with others. These signals can immobilize other cells, kill or eat prey or form a large community.
Microbes secrete specific signals to communicate with others. These signals can immobilize other cells, kill or eat prey or form a large community.- Microbes are able to integrate multiple sources of information—signals related to many different senses such as chemicals, temperature and touch. Using this successfully integrated information, they make decisions as to movement and other behavior.
- When they are not responding to sensory stimuli, microbes act as individuals and respond to internal clocks and other sources of internal information. They can move without sensory stimuli, such as searching for food.
Internal Signaling With Metabolites
Recent research has begun to provide mechanisms for decision making in a cell. Metabolism, once thought to be only a mechanism to provide fuel and structural molecules, has now been shown to be critical parts of information signaling cascades inside cells.
 A previous post noted that T cells and cancer cells use elements of the respiration (Krebs) cycle as crucial signals, which trigger dramatic alterations in cellular function to rapidly produce an army of cells. Complex decision-making in microbes, also, uses a very wide range of metabolic signals including factors that alter gene networks and modulate the production and shapes of proteins. Key molecules in microbe metabolism not only control production of energy, manufacture of amino acids and proteins, and overall metabolism, but also carry specific information, which is a basis of decision-making.
A previous post noted that T cells and cancer cells use elements of the respiration (Krebs) cycle as crucial signals, which trigger dramatic alterations in cellular function to rapidly produce an army of cells. Complex decision-making in microbes, also, uses a very wide range of metabolic signals including factors that alter gene networks and modulate the production and shapes of proteins. Key molecules in microbe metabolism not only control production of energy, manufacture of amino acids and proteins, and overall metabolism, but also carry specific information, which is a basis of decision-making.
Metabolism fuels all activities with energy and materials. Microbes must find food and then must metabolize this matter, making small and large molecules. The microbe transmits information about these energy-producing processes to other cellular functions such as movement and communication. Specific molecular circuits have been identified that gather specific information and make decisions.
 This new view of microbes is based on recent advances in measuring fluctuations of metabolic circuits—concentration of products of metabolism and identification of specific protein amounts and activities. Measurements have been made of the information input that informs microbes of their cellular or external environment conditions, as well as, the output or decisions that are made to maintain or alter the metabolism of the bacteria.
This new view of microbes is based on recent advances in measuring fluctuations of metabolic circuits—concentration of products of metabolism and identification of specific protein amounts and activities. Measurements have been made of the information input that informs microbes of their cellular or external environment conditions, as well as, the output or decisions that are made to maintain or alter the metabolism of the bacteria.
Common molecules that are important in metabolic cycles are also critical information signals—fructose 1,6-biphosphate (FBP), glutamine and ATP (the universal phosphorus energy particle). Although these are universally part of metabolism, the ways they are used as information varies in different microbes.
Multiple Microbe Membrane Receptors
 Recently it has been shown that microbes’ membranes are a vast complex structure made of thousands of receptors. These take the form of a lattice of hexagonal structures that appear as a honeycomb. The receptor network is made up of many different kinds of proteins—some simple receptors that trigger cascades with coupling proteins, and some enzymes. In the diagram on the left receptor molecules are pink, coupling proteins are green and enzymes are blue.
Recently it has been shown that microbes’ membranes are a vast complex structure made of thousands of receptors. These take the form of a lattice of hexagonal structures that appear as a honeycomb. The receptor network is made up of many different kinds of proteins—some simple receptors that trigger cascades with coupling proteins, and some enzymes. In the diagram on the left receptor molecules are pink, coupling proteins are green and enzymes are blue.
New types of receptors are being discovered frequently. One example is in B. mycoides, which has a sense of touch and can respond to the shape and curvature of its surroundings. Individual microbes respond by producing curving filaments that has a totality forms spiral art.
Receptors for Carbon Food Sources Trigger Different Mechanisms
 Microbes love carbon food sources and have many receptors related to carbon for different situations. To save energy, they only produce receptors and special transporters as needed, based on signals from within the cell and the outside environment. There are, also, membrane receptors for phosphate, nitrate, sugars, and thirty more external substances.
Microbes love carbon food sources and have many receptors related to carbon for different situations. To save energy, they only produce receptors and special transporters as needed, based on signals from within the cell and the outside environment. There are, also, membrane receptors for phosphate, nitrate, sugars, and thirty more external substances.
Several mechanisms are utilized to gather the information from the many different membrane receptors. One type involves two molecules—one in the membrane and another connected just below it. Another type serves as a single receptor and transporter. Both trigger cascades of enzymes to gather the information internally.
A different receptor, for specific carbon sources, stimulates enzymes that both sense the signal and respond themselves to increase specific pathways that can consume that particular type of carbon food. These single internal sensors are the predominant mechanism for many food sources, such as glucosamine, fucose, maltose and many others. A signal in this system makes a small adjustment to increase detection of this particular food, but at the same time it signals to the gene networks in the nucleus to make large adjustments. The metabolic intermediary molecule accumulates, which then signals to the nucleus and triggers a bigger response to upregulate the entire process.
Multiple Sources at Once
 The simple feedback loop just mentioned does not choose priorities when multiple carbon sources occur at the same time. In a more advanced mechanism, several sources are recognized and the preferred sources are consumed first with upregulation of that specific mechanism. When exhausted, the next nutrient cycle is revved up through specific signals.
The simple feedback loop just mentioned does not choose priorities when multiple carbon sources occur at the same time. In a more advanced mechanism, several sources are recognized and the preferred sources are consumed first with upregulation of that specific mechanism. When exhausted, the next nutrient cycle is revved up through specific signals.
Circuits notice the better source and inhibits the other cycles. They, also, keep track of the priorities. One example is when E. coli takes up glucose—its most preferred nutrient. When a molecule in that cycle is altered (EIIA is dephosphorylated) it becomes a signal that inhibits transporters for several other sources of carbon.
But, E. coli, also, uses another system to make sure that the transport and metabolism are inhibited for the other receptors. This signal, Crp, emphasizes several of the most preferred food sources. It is activated by cyclic AMP and by the original phosphorylated EIIA.
Another circuit uses pyruvate and other prominent molecules that use amino acids as its signal. These metabolites trigger the Crp pathway relative to the ratio of the carbon breakdown and synthesis activities. It stimulates genetic networks in one direction when there is more carbon than nitrogen, and the other direction with the opposite ratio.
Central Carbon Metabolism Circuits
 Because of the multiple overlapping complex cycles, it is much more difficult to study central carbon metabolism than the simpler mechanisms just described for external nutrients. In the fast moving central metabolism, altering genetic networks is too slow and ineffective. Instead, the regulation uses alterations of proteins through addition of phosphate and acetyl groups that change the binding properties of proteins or specific interactions with other molecules that change the protein’s shape. This protein alteration occurs in seconds, rather than genetic stimulation in minutes.
Because of the multiple overlapping complex cycles, it is much more difficult to study central carbon metabolism than the simpler mechanisms just described for external nutrients. In the fast moving central metabolism, altering genetic networks is too slow and ineffective. Instead, the regulation uses alterations of proteins through addition of phosphate and acetyl groups that change the binding properties of proteins or specific interactions with other molecules that change the protein’s shape. This protein alteration occurs in seconds, rather than genetic stimulation in minutes.
An important example is a switch that operates the fundamental breakdown of sugars to pyruvate, producing energy in the form of ATP. This cycle both blocks the pathway that makes new glucose and inhibits the sugar breakdown. When there is no glucose inside the cell, the pathway makes sure that it is ready for new glucose as it appears from the environment.
Many Overlapping Circuits
 Several specific uses of information have been found in the very complex central metabolic pathways, but there are hundreds of other pathways that are just being studied and are not yet understood.
Several specific uses of information have been found in the very complex central metabolic pathways, but there are hundreds of other pathways that are just being studied and are not yet understood.
The pattern of molecules in the middle of cycles for synthesis and breakdown carries information to other circuits for other purposes. One example (of many) is pyruvate that alters the shapes of several enzymes and two factors that stimulate gene networks (transcription factors). It not only regulates its own consumption, but also is critical in genes related to cell division and many other seemingly unrelated circuits.
Energy Metabolism Circuits
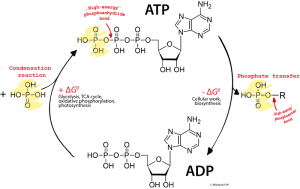
A phosphorus molecule with three attached oxygen atoms and additional hydrogen is called a phosphate and is used as an energy source in processes that take one of the phosphates and use it to trigger chemical reactions. The larger molecule with three phosphates is called ATP—adenosine tri (3) phosphate—and the smaller ADP—adenosine di (2) phosphate. There is, also, a mono adenosine, AMP with even less energy.
The central energy of life uses the addition and subtraction of these phosphates to give and take energy. Two processes that add phosphates and create ATP are respiration and glycolysis (the breakdown of sugar).
In respiration, electrons are taken from carbon and in a complex process they alter oxygen, nitrogen or sulphur. Respiration is the greatest source of ATP but needs to use many proteins. So, if the cell is stressed it can try to increase respiration (the circuit Crp mentioned above) or switch to the less efficient process of the breakdown of sugar, or glycolysis. In this latter process, ATP (3 phosphates), ADP (2 phosphates) and AMP (1 phosphate) are all part of information circuits that regulate increase of different pathways through many different enzymes.
Respiration Cycle Is A Central Information Pathway
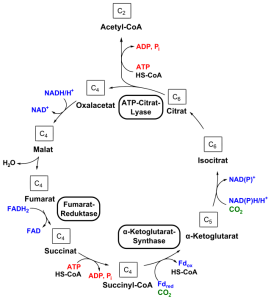 Respiration is called the citric acid cycle, the tricarboxylic acid cycle (TCA cycle), or the Krebs cycle. It is a very complex process that needs many large proteins for the electron transfer. Regulation of respiration occurs prominently through stimulating gene networks that make specific proteins. It is, also, an expensive pathway that needs many more ribosomes (big complex molecules). In cells that are growing, it is more economical to reduce the need for respiration.
Respiration is called the citric acid cycle, the tricarboxylic acid cycle (TCA cycle), or the Krebs cycle. It is a very complex process that needs many large proteins for the electron transfer. Regulation of respiration occurs prominently through stimulating gene networks that make specific proteins. It is, also, an expensive pathway that needs many more ribosomes (big complex molecules). In cells that are growing, it is more economical to reduce the need for respiration.
One factor senses the amount of oxygen available in the cell and can stop respiration genes and start pathways that don’t use oxygen. Another factor measures the respiratory molecules that appear when oxygen is low, which then stimulates another pathway. E. coli uses three mechanisms to regulate the respiration cycle—carbon, oxygen and energy. These three signals are then integrated
Nitrogen Pathways
 The pathways to measure and regulate nitrogen uptake and metabolism are very similar to carbon. Ammonium is the preferred source that is prioritized over others. But, there are, in fact, many different sources of nitrogen in the environment. The cell detects the transporters of nitrogen and then revs up the needed pathway molecules.
The pathways to measure and regulate nitrogen uptake and metabolism are very similar to carbon. Ammonium is the preferred source that is prioritized over others. But, there are, in fact, many different sources of nitrogen in the environment. The cell detects the transporters of nitrogen and then revs up the needed pathway molecules.
E. coli has a very complex information system to understand the availability of nitrogen—similar to the Crp system of carbon. Complex circuits are stimulated when there is less available nitrogen. The information signal is glutamine concentration, a critical molecule that supplies nitrogen for reactions. Glutamine alters several factors that stimulate different pathways. Glutamine pathways are inhibited by many different molecules that are produced by nitrogen metabolism—glutamine itself, amino acids, and nucleotides.
Several molecules from the carbon cycles are, also, used with nitrogen that maintains the balance between them. Glutamine, also, stimulates gene factors that are related to specific outside sources of nitrogen. What is interesting is that these circuits are really unrelated but maintain one signal for all of them. Glutamine is, also, able to suppress non-preferred outside sources.
 Individual amino acids use a variety of pathways for manufacture and breakdown. Each of the twenty amino acids in E. coli has its own regulatory network, where the concentration of the amino acid determines how much is needed and revs up the production or inhibits it. The inhibition occurs in the first step of synthesis through altering the enzyme shape, which is a rapid response. Each amino acid is regulated independently of the others. Ten of the amino acids use gene transcription factors to inhibit their own synthesis circuits. They, also, have special signals to break up existing supplies when there is too much.
Individual amino acids use a variety of pathways for manufacture and breakdown. Each of the twenty amino acids in E. coli has its own regulatory network, where the concentration of the amino acid determines how much is needed and revs up the production or inhibits it. The inhibition occurs in the first step of synthesis through altering the enzyme shape, which is a rapid response. Each amino acid is regulated independently of the others. Ten of the amino acids use gene transcription factors to inhibit their own synthesis circuits. They, also, have special signals to break up existing supplies when there is too much.
Even though the amino acid pathways are separate, there are, also, overriding pathways that affect all of them. The amino acid leucine, for unknown reasons, is a master regulator and stimulates hundreds of gene networks regulating manufacture of less or more amino acids. It can shut down the entire process during a phase without growth or starvation. This network operates in decision-making pathways related to the poles of growth and starvation.
Protein Pathways
 When cells are growing rapidly, most energy is utilized for making proteins and producing more ribosomes, the factories that manufacture proteins. Surprisingly, 75% of all genetic production can be consumed making the necessary ribosomes.
When cells are growing rapidly, most energy is utilized for making proteins and producing more ribosomes, the factories that manufacture proteins. Surprisingly, 75% of all genetic production can be consumed making the necessary ribosomes.
Deciding whether there is enough material to manufacture a complex molecule like a protein requires integration of a many different information pathways. Two major sources of this ribosome regulation are the amount of available energy (ATP) and amino acids. There are many different pathways that make sure when ribosomes are made or inhibited. These pathways utilize both slow (genetic) and fast (alterations of enzyme shape) mechanisms.
Regulation of proteins and ribosomes is very complex, because in any pathway, including stress pathways, many enzymes are needed. Given that the total production of all proteins is regulated, the cell, then, must balance specific protein production with the necessary ribosomes. Rapidly growing cells switch from respiration to the less protein intensive sugar breakdown cycle to save energy. Prioritizing specific enzyme production with enough ribosomes involves integrating many different signals.
How Do Microbes Make Decisions

The regulation of even a small microbe involves hundreds, or thousands, of different one step and two-step receptors, information particles, regulators, modulators and molecular cascades. These processes use genetic stimulation and alteration of protein shapes. Concentrations of different molecules are signals for different circuits. Most of the factors in E. coli that bind to genes are related to information molecules in metabolism cycles.
Microbes respond to hundreds of receptors, and a large number of internal signals providing information that enables the cell to respond to a huge range of environmental conditions, as well as, the signaling to other microbes for group actions. One view is that the cell uses rapid protein altering mechanisms to regulate materials and to fix gross imbalances. Slower genetic mechanisms respond to major changes in the environment.
The cell integrates local and global signals and cycles. This includes not only understanding the amount of carbon and other nutrients available in the environment, but also exactly what the cell needs throughout all of the hundreds of different pathways and responses. This means that many different loops interact between global information setting parameters and local loops for each of the factors involved. The largest is the cell’s total capacity related to supply and demand of protein and ribosome production and total available energy.
 This new research is giving a glimpse of the microbe metabolic “brain.” But, teasing out the hundreds of interacting factors, circuits, cycles and loops is tremendously difficult. It is similar to the work of deciphering the network of brain cells, only much smaller. The current discussion is based on what has been recently learned, but there is an enormous amount that is unknown.
This new research is giving a glimpse of the microbe metabolic “brain.” But, teasing out the hundreds of interacting factors, circuits, cycles and loops is tremendously difficult. It is similar to the work of deciphering the network of brain cells, only much smaller. The current discussion is based on what has been recently learned, but there is an enormous amount that is unknown.
We are still a very long way from understanding the extremely complex specific microbe communication and the resultant behavioral changes. The cellular intelligence needed for such complex communication that alters behavior is quite startling.