50% of the human genome consists of jumping genes or mobile genetic elements.
50% of the human genome consists of jumping genes or mobile genetic elements.
The 8% of human DNA from retroviruses has been vital to human evolution, such as determining the human placenta, epigenetic changes in the brain and digestive enzymes.
An epigenetic immune system in the nucleus battles the jumping genes for control of the cell and control of evolution.
Jumping genes, being large strands of DNA with specific functions, are much more likely to be the drivers of evolution than simple mutations.
It is not possible to distinguish the many virus-like particles and jumping genes.
Now, Polintons are a new link between viruses, jumping genes, and evolution
 Recently, many new types of virus-like particles, or mobile genetic elements, have been found. They are in the three classes of cells—archaea, bacteria, and eukaryotes. Ubiquitous transfer of genetic material between all of these cells blurs any classification as well as the evolution of all of these cells. Many different genetic particles can live in cells and greatly influence them. They can become part of the cell’s genomes; produce viruses used to attack other cells and become jumping genes. Retroviruses are a well-known mobile genetic element that uses messenger RNA to make DNA, which is inserted into the human genome. Epigenetic tags of DNA methylation silence retroviruses and jumping genes. Recently, it was discovered retroviruses play a very active role in brain cells. Now, a link between retroviruses, jumping genes and other genetic mobile elements is found in the newly discovered Polinton jumping genes and viruses. Polintons are a vital link in the evolution of intelligent viruses and jumping genes
Recently, many new types of virus-like particles, or mobile genetic elements, have been found. They are in the three classes of cells—archaea, bacteria, and eukaryotes. Ubiquitous transfer of genetic material between all of these cells blurs any classification as well as the evolution of all of these cells. Many different genetic particles can live in cells and greatly influence them. They can become part of the cell’s genomes; produce viruses used to attack other cells and become jumping genes. Retroviruses are a well-known mobile genetic element that uses messenger RNA to make DNA, which is inserted into the human genome. Epigenetic tags of DNA methylation silence retroviruses and jumping genes. Recently, it was discovered retroviruses play a very active role in brain cells. Now, a link between retroviruses, jumping genes and other genetic mobile elements is found in the newly discovered Polinton jumping genes and viruses. Polintons are a vital link in the evolution of intelligent viruses and jumping genes
The little known Polintons started at the dawn of eukaryote evolution and are at the center of the evolution of many very important viruses and jumping genes. They became human viruses, transposons (jumping genes) and plasmids. They exist in mitochondria as linear plasmids, as well as eukaryote viruses. Polintons could, also, be the link between the mitochondrial ancestors and the eukaryote cell ancestors.
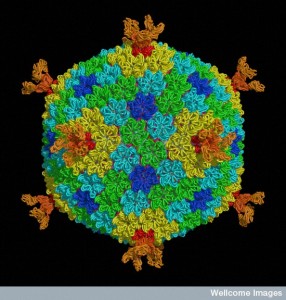 Because of the constant horizontal exchange of genes, it is very difficult to understand how viruses and virus particles, such as jumping genes, evolved. A new breakthrough is finding a virus family connected with a jumping gene family, called Polintons. Polintons are very closely related to very large viruses and very small viruses that feed on larger viruses. They are, also, closely related to the virus phages that feed on, and are satellites and tools of, bacteria. They connect all of these to jumping genes and many other small mobile elements.
Because of the constant horizontal exchange of genes, it is very difficult to understand how viruses and virus particles, such as jumping genes, evolved. A new breakthrough is finding a virus family connected with a jumping gene family, called Polintons. Polintons are very closely related to very large viruses and very small viruses that feed on larger viruses. They are, also, closely related to the virus phages that feed on, and are satellites and tools of, bacteria. They connect all of these to jumping genes and many other small mobile elements.
Virus Evolution
There are a vast amount of different viruses and many different mobile virus-like particles, with many different structures, mechanisms of survival and ways that cellular genes are manipulated.
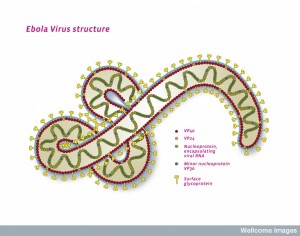 Viruses are so diverse that there is no single gene that is in all of them. In fact, they can be extremely effective with only 7 viruses and nine proteins, such as Ebola. The many different kinds of mobile elements, however, have developed relationships among themselves, which come from the ubiquitous transferring of genetic material in all cells.
Viruses are so diverse that there is no single gene that is in all of them. In fact, they can be extremely effective with only 7 viruses and nine proteins, such as Ebola. The many different kinds of mobile elements, however, have developed relationships among themselves, which come from the ubiquitous transferring of genetic material in all cells.
As genes evolve they are shared among many different kinds of cells and are used in varied ways. A well-known example is the use of viral genes in the human genome to make the human placenta (see post). The most important genes for viruses are the ones that help the virus to copy itself. And these critical genes are copied and transferred widely.
Different viruses attack the three basic types of living cells—eukaryotes, bacteria and archaea. Retroviruses, which have developed or rather stolen, the ability to place themselves into the human genome, exist only in eukaryotes.
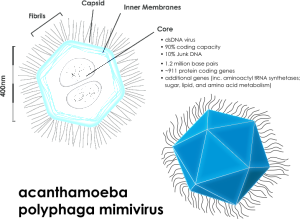
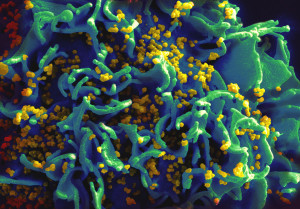 Eukaryotes with much more complex and larger cells have many different types of transposable elements including a wide variety of RNA viruses, which can live with fewer enzymes and move more quickly. New viruses are being discovered frequently, such as the recent discoveries of the very large Mimivirus and Pandoravirus (please see post on Giant Viruses in Evolution.) These large viruses have their own smaller viruses called either virophages or satellites. Satellites are dependent on the larger virus; virophages influence the large virus by decreasing its function and keepin the host cell alive longer..
Eukaryotes with much more complex and larger cells have many different types of transposable elements including a wide variety of RNA viruses, which can live with fewer enzymes and move more quickly. New viruses are being discovered frequently, such as the recent discoveries of the very large Mimivirus and Pandoravirus (please see post on Giant Viruses in Evolution.) These large viruses have their own smaller viruses called either virophages or satellites. Satellites are dependent on the larger virus; virophages influence the large virus by decreasing its function and keepin the host cell alive longer..
May Kinds of Virus-Like Particles or Mobile Genetic Elements
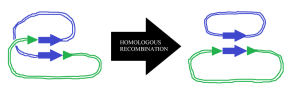
There are many ways pieces of DNA and RNA are transferred between cells. These are called either horizontal gene transfer (HGT) or transfer of mobile elements. Each strand of DNA or RNA can transfer unique functions in the process that can be used in many different ways in evolution. Types of mobile elements are:

From Famedog Human transposons cut and paste their way in and out of human DNA in three ways.
- Long terminal repeats (LTR) use the retrovirus enzyme.
- Long interspersed elements, LINE, use a different enzyme.
- Short interspersed elements, SINE, uses an RNA enzyme.
- Retrotransposons are copied in two stages, transcribed from DNA to RNA, then reverse transcribed back to DNA in a different spot. This second step is similar to retrovirus using the same enzyme.
- Retroviruses are transposable elements like jumping genes.
- Satellite viruses, are dependent on large viruses for their function.
 Virophages, like Sputnik and Mavirus are very small viruses that colonize and influence larger viruses. Mavirus shares 7 proteins and most genes with the Polinton jumping gene. They usually diminish the numbers and role of the larger virus in the host cell and keep the cell alive longer.
Virophages, like Sputnik and Mavirus are very small viruses that colonize and influence larger viruses. Mavirus shares 7 proteins and most genes with the Polinton jumping gene. They usually diminish the numbers and role of the larger virus in the host cell and keep the cell alive longer.- Transpovirons are small retroviruses for other viruses. They can insert themselves anywhere into the genes of a giant virus.
- Viroids are genetic elements in plants.
- Plasmids are a small, often circular, piece of DNA that exists separate from the chromosome DNA in a cell and is transmitted from one cell to another by horizontal gene transfer HGT. They can function independently inside the cell. Plasmids can carry antibiotic resistance genes among.
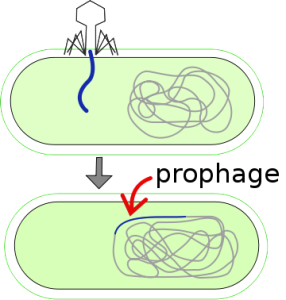
 Prophage is the DNA of a phage virus that is inserted into the circular bacterial DNA chromosome or as an independent plasmid. This is a latent bacteriophage and may be used by bacteria to “create” a virus to fight enemies.
Prophage is the DNA of a phage virus that is inserted into the circular bacterial DNA chromosome or as an independent plasmid. This is a latent bacteriophage and may be used by bacteria to “create” a virus to fight enemies.- Self-splicing introns are ribozymes are RNA based molecules that function as enzyme catalysts for reactions. These particles are mobile, are transmitted between cells and are active in self-splicing and self-editing of all kinds.
- Inteins are a piece of a protein that acts like a virus or jumping gene by cutting and sewing itself to join parts of the protein called This occurs through protein splicing, not DNA splicing.
- Insertion sequences (IS) are short piece of DNA that act like a jumping gene.
- Self-editing alternative RNA transcripts are alternative RNA transcripts or self directed RNA that are critical to brain evolution. (see post RNAs in brain evolution.
In addition three new transfer mechanisms have been discovered.

From Ayacop Tunneling nanotubes are long tubes between cells to exchange genetic material. They were only discovered recently because they are so small, but appear to exist between most cells.
- Exosomes are sacs made from membranes that send genetic material between cells. This, also, appears to be extremely common.
- Dodder plant has haustoria that suck up genetic material from a plant and then transfers it to other plants.
Polintons At the Core of Jumping Genes and Human Viruses
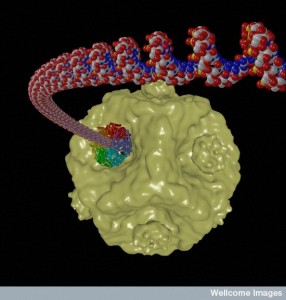 Polintons, or Mavericks, are a unique form of very ancient transposon jumping gene in vertebrates, including humans, made of large pieces of double stranded DNA. They are very closely related to Mavirus virophages, which have 19K DNA bases and 20 proteins. The Polinton jumping gene has 9K to 22K DNA bases and shares many proteins with Mavirus virophage. Each of these large jumping genes produces two different proteins that are similar to those that cover viruses (capsids). They can, also, produce mobile virus-like particles that have many different behaviors. Polintons bridge the two major types of mobile elements—jumping genes and viruses.
Polintons, or Mavericks, are a unique form of very ancient transposon jumping gene in vertebrates, including humans, made of large pieces of double stranded DNA. They are very closely related to Mavirus virophages, which have 19K DNA bases and 20 proteins. The Polinton jumping gene has 9K to 22K DNA bases and shares many proteins with Mavirus virophage. Each of these large jumping genes produces two different proteins that are similar to those that cover viruses (capsids). They can, also, produce mobile virus-like particles that have many different behaviors. Polintons bridge the two major types of mobile elements—jumping genes and viruses.
Polintons make many different versions of critical protein enzymes common in viruses. These include critical DNA polymerase that copies the genetic material; integrase that sews DNA in and out of genomes; protease that cuts proteins; ATPase that grabs high-energy phosphate particles to use for energy; and several proteins for the capsid cover. Some Polintons have other proteins found in viruses and jumping genes, such as helicases and primases. These form a complex with polymerase involved in synthesizing RNA and DNA.
Among all of the shared genes in different viruses and transposons, it appears that Polintons are at the center of many different species. A large number of genes are shared between Polintons, plasmids, transposons, major virus families, and virophages. Polintons appear to be the hub of a wide variety of viruses, plasmids and jumping genes.
Polinton Proteins
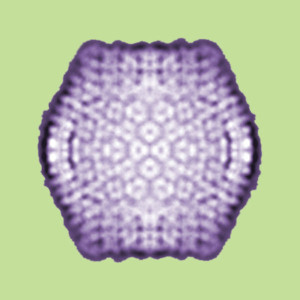
 Polintons make proteins that are similar to the Major and Minor Capsid Proteins. These proteins construct the complex icosahedral geometric forms of the common double stranded DNA virus covers for all three cellular kingdoms. Many viruses add other structural proteins to make more varied designs.
Polintons make proteins that are similar to the Major and Minor Capsid Proteins. These proteins construct the complex icosahedral geometric forms of the common double stranded DNA virus covers for all three cellular kingdoms. Many viruses add other structural proteins to make more varied designs.
The Polinton major capsid protein is similar to the large Megavirales family of viruses. This connects Polintons with the largest viruses and with smaller double stranded viruses.
 Minor capsid proteins make five pointed structures for many virus covers. These are in a many different viruses, but not the very large viruses. They do appear in some large viruses such as Mimi, but not the giants. The genetic arrangement is similar to the Mavirus (Maverick).
Minor capsid proteins make five pointed structures for many virus covers. These are in a many different viruses, but not the very large viruses. They do appear in some large viruses such as Mimi, but not the giants. The genetic arrangement is similar to the Mavirus (Maverick).
Adenoviruses are important human viruses. A critical cysteine protease in adenovirus is from Polintons; it is needed to release the final infectious virus out of a cell. This protein, also, occurs in Megaviruses. The protease seems to be a very ancient part of the eukaryote and it is not in Achaea or bacteria (no proteases). Human transposons, also, have the same integrase and the protease proteins.
This protease is one big difference between Polintons and bacterial viruses.
Polinton Evolution
Politons followed two different paths based on their ability to live in the nucleus and the genome. One was as a transposable element in vertebrate cells with the ability to sew itself in and out of the genome. The other is as a virus. In the genome they could live for a long time and gradually gather alterations from recombination without the pressure of virus life, such as having to build new viruses constantly and adapt to the cell’s evolving combat (see post on War between Microbes and Cells). Some of the jumping genes could wait quite a long time and then “jump” out of the genome to become a virus again.
The Polinton integrase and protease give them the ability to sew themselves into the DNA genes of the vertebrates making them very successful jumping genes. They can live in the genome and produce virus like particles, jumping genes, or full-blown viruses. One group of Polintons became committed to being a jumping gene and gave up genes needed for virus life. They became up to 30% of a cell’s genome.
The Polinton enzyme that uses energy to package the virus inside the covering capsule is similar to a variety of different viruses showing the wide range of gene transfer that has occurred through Polintons.
 A particular type of DNA replication style called “protein prime” is from the Polinton line. This type of copying of DNA is started with a special protein that attaches at a particular point (5’ end) to the genes. It is unique and different from the type that cells and many other viruses use—nucleic acid to mark the beginning of the copying. There are many variations of the terminal protein that are used for the “protein priming”.
A particular type of DNA replication style called “protein prime” is from the Polinton line. This type of copying of DNA is started with a special protein that attaches at a particular point (5’ end) to the genes. It is unique and different from the type that cells and many other viruses use—nucleic acid to mark the beginning of the copying. There are many variations of the terminal protein that are used for the “protein priming”.
Research about the Polinton polymerase shows evolution of two separate lines. One is among bacteria and archaea. The other is with viruses of eukaryotes and plasmids.
The eukaryote type has two types, as well. One is in mitochondria of plants and fungus with a DNA polymerase, an attached terminal protein, and another RNA polymerase. The second is in plasmids in yeast of eukaryote cells with protein priming. This is directly from the Polintons and is related to many other viruses.
Many Species From Polintons
 The structures of the human adenovirus capsids and bacteria and archaea tectiviruses capsids are identical and both are from Polintons. Another evolutionary twist was that by adenoviruses losing integrase, they became very active mobile viruses that don’t go into the genome like retroviruses. Polintons are the link between these two important viruses for humans and for bacteria. Also, the important four types of adenoviruses evolved late in vertebrate evolution by transfer of material from Polintons.
The structures of the human adenovirus capsids and bacteria and archaea tectiviruses capsids are identical and both are from Polintons. Another evolutionary twist was that by adenoviruses losing integrase, they became very active mobile viruses that don’t go into the genome like retroviruses. Polintons are the link between these two important viruses for humans and for bacteria. Also, the important four types of adenoviruses evolved late in vertebrate evolution by transfer of material from Polintons.
Bidnavirus insect viruses have the same polymerase from the Polintons. This occurred by multiple genetic exchanges from Polintons infecting insects.
Virophages and satellite viruses that are similar to Mavirus came with the transfers of polymerase from Polintons. These greatly confuse any evolutionary tree because of their great diversity. The virophage sputnik related to the large Mimivirus has a very diverse set of polymerases. Sputnik has been responsible for spreading many genes in evolution.
 Some yeasts have plasmids that stay outside of the nucleus and carry their own replication proteins. These, also, are derived from Polintons, and are similar to adenovirus in humans. The important human poxviruses come from these plasmids. Linear plasmids come from Polintons, the adenovirus ancestors. An old Polinton somehow left the nucleus with the new proteins and then evolved into Magavirales and these plasmids in the cytoplasm.
Some yeasts have plasmids that stay outside of the nucleus and carry their own replication proteins. These, also, are derived from Polintons, and are similar to adenovirus in humans. The important human poxviruses come from these plasmids. Linear plasmids come from Polintons, the adenovirus ancestors. An old Polinton somehow left the nucleus with the new proteins and then evolved into Magavirales and these plasmids in the cytoplasm.
From Polintons, Megavirales was given the vital genes to build a virus, except they developed their own transcription enzyme so they didn’t have to stay in the nucleus and use the cell’s machinery. This line produced plasmids in a simpler lifestyle as well as Megavirales in a much more complex lifestyle. The polymerase changed to allow bigger viruses.
Viruses, Polintons and Jumping Genes in Evolution
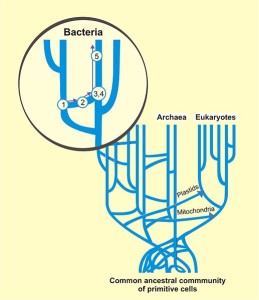 Each human has ten trillion cells, hundreds of trillion bacteria and billions of trillion viruses. Only a very small number of microbes are understood. The total amount of microbe genetic material in humans is 300x the total human genome and many greatly influence humans. This total amount of functional DNA and RNA is called the hologenome.
Each human has ten trillion cells, hundreds of trillion bacteria and billions of trillion viruses. Only a very small number of microbes are understood. The total amount of microbe genetic material in humans is 300x the total human genome and many greatly influence humans. This total amount of functional DNA and RNA is called the hologenome.
Viruses and virus like particles are constantly transferring genetic material between all cells, and therefore the “tree of life” is not really understood. Viruses are, however, critical in all branches of the tree because of the constant genetic transfer. There are, also, many different virus-like particles at work.
Recent research (ENCODE – encyclopedia of DNA elements – see post) shows that a large amount of the DNA besides the 1.5% known to be human genes are probably active and not “junk”. They make relevant and usable RNAs of all types, large and small, that have many varied functions. But, 50% of the DNA consists of jumping genes, retroviruses and other mobile elements.
Virus DNA produces critical promoters and enhancers that alter human gene function and affect alternative RNA splicing. Many cancers are caused by retroviruses. Salivary amylase, allowing humans to eat starch, comes from virus DNA. They are essential for fetal stem cells, the placenta and mother’s immune system.
Evolution is now known to be critically dependent on the ubiquitous gene transfer between all cells, which become jumping genes and retroviruses. Epigenetics has evolved in defense of these 50% jumping genes in the nucleus. See other posts for a detailed description of the epigenetic immune system that battle jumping genes in the nucleus (CRISPR – post giant viruses).
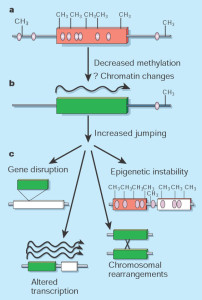 Bacteria minimize the amount of DNA they use; they give up functions to others in order to travel light. Prokaryotes use the same regulatory mechanisms to control them, but jumping genes don’t proliferated in prokaryotes.
Bacteria minimize the amount of DNA they use; they give up functions to others in order to travel light. Prokaryotes use the same regulatory mechanisms to control them, but jumping genes don’t proliferated in prokaryotes.
Eukaryotes, in contrast, accumulate large amounts of extra jumping genes, both copied and altered. One of the major differences between eukaryotes and prokaryotes is the more elaborate epigenetic mechanisms in eukaryotes from this battle with mobile genetic elements. The mobile elements give the cell much more material to work with for complex evolution, but, also, demands much more regulation. The regulation includes regulatory proteins, regulatory RNA, methylation of DNA and of acetylation of histones. These mechanisms have evolved along with the huge proliferation of DNA elements from jumping genes and viruses. Epigenetic mechanisms are not only critical for the entire transcription process and structure, but they stimulate more jumping genes.
These two opposing forces of mobile elements and epigenetics are very related—the proliferation of mobile DNA particles and the cell’s epigenetic mechanisms to control them. With environmental stress, the genome responds with the activation of the epigenetic processes. This increases the capacity to withstand this stress, which can be inherited through epigenetic mechanisms. When the stress subsides then the increased amount of jumping gene activity also subsides.
Evolution of Intelligent Viruses and Jumping Genes
With constant transfer of the many different mobile genetic particles, the entire understanding of evolution has changed. Viruses, virus satellites, virophages and plasmids are responsible for transferring large strands of DNA that have many different specific functions. In the new home, these functions adapt into new functions. This is responsible for the spread of antibiotic resistance, but, also, the human placenta, digestive enzymes and many cancers.
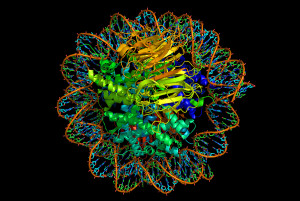 Once placed in the human genome, these genetic particles become jumping genes that make up almost 50% of the human genome (retroviruses are 8%). Human cells have welcomed these jumping genes as drivers of evolution, but, also, constantly battle them through elaborate epigenetic mechanisms. In fact, in evolution, jumping genes, viruses and epigenetics developed in relation to each other.
Once placed in the human genome, these genetic particles become jumping genes that make up almost 50% of the human genome (retroviruses are 8%). Human cells have welcomed these jumping genes as drivers of evolution, but, also, constantly battle them through elaborate epigenetic mechanisms. In fact, in evolution, jumping genes, viruses and epigenetics developed in relation to each other.
Now, an ancestor of the major viruses, and virus particles, as well as the human jumping genes has been found. These Polintons are at the center of many very important aspects of evolution.
Previous posts have described the incredible intelligent behavior of viruses (Ebola, HIV, Varicella). Jumping genes demonstrate this same intelligent activity. These mobile elements have stimulated the large eukaryote cell to defend itself. In fact, it is intelligent warfare between mobile genetic elements and the epigenetic responses of the cell that, most likely, determines evolution.